|
APOBEC-1 editing of apolipoprotein B mRNA
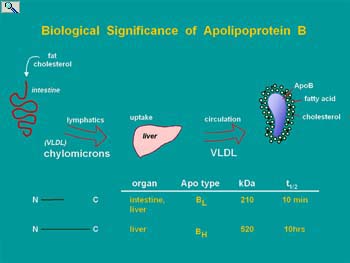
The post-transcriptional editing of mRNA encoding apolipoprotein B (apoB) involves deamination of cytidine at nucleotide position 6666 to form uridine. The resultant conversion of a CAA glutamine codon to a translation stop UAA codon enables two proteins with different functional properties to be encoded from one gene. ApoB is a non exchangeable structural component of intestinal-derive chylomicrons and hepatic derived very low density (VLDL) and low density (LDL) lipoprotein particles. Hepatic apoB mRNA editing is a highly regulated process that determines the proportion of VLDL that contain full length (apoB100) or truncated (apoB48). As only apoB100 VLDL are metabolized in the blood to the LDL ( a known atherogenic risk factor associated with cardio-vascular disease and stroke), editing and mechanisms regulating the abundance of edited apoB mRNA in liver are of biomedical significance.
|
|
|
ApoB protein transports lipid in the blood stream. Lipase in the serum will digest the triglycerides of VLDL converting them to cholesterol and protein rich LDL, a risk factor for atherosclerosis. This takes time and only occurs on VLDL assembled on apoB100. ApoB48 containing VLDL (arising from edited apoB mRNA) are cleared from the blood too rapidly to be converted to LDL and hence have the potential of transporting lipid without as much risk of contributing to disease.
|
|
|
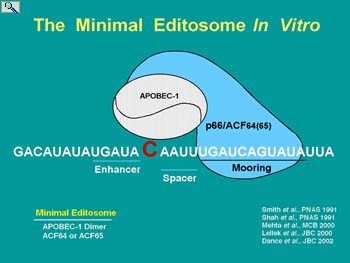
APOBEC-1 is the cytidine deaminase responsible for the apoB mRNA editing but its capacity to do so is dependent on interactions with RNA binding auxiliary proteins known collectively as APOBEC-1 Complementation Factor, ACF. Our research addresses recent observations suggesting the hypothesis that insulin regulation of editing activity occurs at multiple levels including APOBEC-1 gene transcription, ACF and APOBEC-1 trafficking between the cytoplasm and the nucleus (the site of editing), alternative splicing of mRNAs encoding ACF and post-translation modification of editing factors. Moreover, the abundance of edited apoB mRNA for translation is regulated by ACF-dependent resistance to nonsense mediated RNA decay (NMD). The goals of this research are to demonstrate what role each spliced ACF variant plays in the metabolic regulation of hepatic editing at the level of their expression, post-translational modification and intracellular trafficking.
|

|
The significance of this research is that it will provide an understanding of the factors involved in regulating high fidelity apoB mRNA editing that may used for the induction of apoB mRNA editing in human liver for therapeutic intervention in atherogenic diseases. Knowledge of how auxiliary proteins interact with editing enzymes to control… site-specific activity will facilitate studies on other related enzymes (AID, APOBEC3G) with roles in controlling immunoglobulin expression and inhibiting retroviral infectivity.
|

|
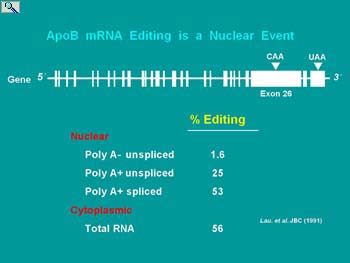
ApoB pre-mRNA is encoded with numerous exons that have to be spliced in order to create a mature mRNA. Editing occurs at position 6666 (CAA) within exon 26. This introduces a premature stop codon up stream of the native stop codon within the terminal exon (UAA). For reasons that are not totally clear but appear to involve ACF binding to the mooring sequence, edited apoB mRNA is not subjected to nonsense mediated decay (NMD).
ApoB mRNA editing was first suggested to be a nuclear process from the analysis of the proportion of edited apoB mRNA among intermediates in the biosynthesis of processed nuclear mRNA and cytoplasmic apoB mRNA. The proportion of apoB mRNA molecules that will be edited, have been edited by the time nuclear mRNA is polyadenylated and spliced.
|
|
|
|
|
|
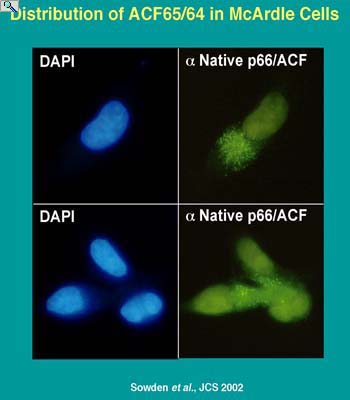
Trafficking of editing factors is integral to the regulation of editing activity. Both APOBEC-1 and ACF are predominantly localized in the cytoplasm of resting liver cells. Hormonal stimulation of apoB mRNA editing activity is coincident with nuclear import of ACF (native ACF shown in green) to the nucleus (DAPI stained blue). It is currently unknown whether ACF transcription is regulated but alternative splicing of ACF premRNA is metabolically controlled. APOBEC-1 transcription is regulated as well as its trafficking between the nucleus and the cytoplasm. The domains of ACF and APOBEC-1 required for trafficking are the subject of investigation.
|
|
|
The lab has also shown a dose-dependent increase in apoB mRNA editing when rat primary hepatocytes are exposed to ethanol. Molecular analysis of apoB mRNA demonstrated that the RNA editing response to ethanol occurred rapidly (within minutes to 2 h) and was sustained (7-18 h). The hypothesis suggested by the data is that the effect of ethanol on RNA editing resulted from cell signaling processes leading trafficking of editing factors and an increase in the assembly of functional editosomes. This process occurred in the absence of de novo RNA or protein synthesis suggesting that the function of pre-existing editing factors can be modulated.
The goal of this branch of our research is to characterize the macromolecules involved in transducing the ethanol signal and to delineate the molecular targets that ethanol might be used to activate trafficking of editing factors and assembly of editosomes.
|
|
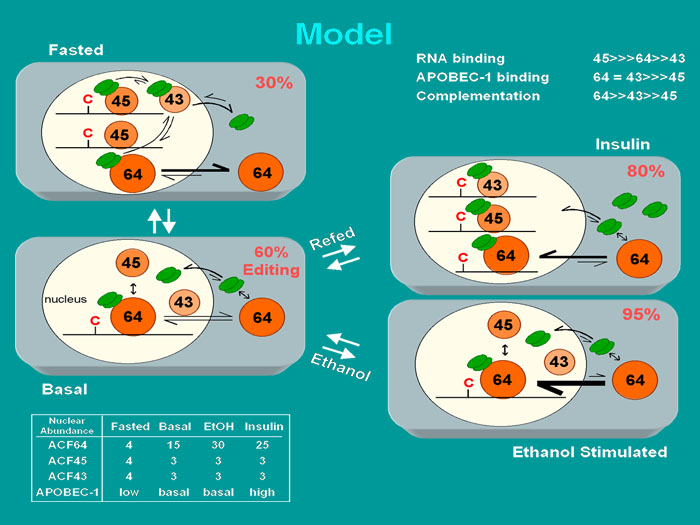
Model for the regulation and catalytic turnover of the editosome. Insulin and ethanol both stimulate apoB mRNA editing whereas fasting (reduced insulin) inhibits editing. Trafficking of the auxiliary proteins ACF64 and APOBEC-1 occurs under metabolic modulation of editing activity . In addition insulin uniquely induces transcription of the APOBEC-1 gene and increases APOBEC-1 protein abundance. Percentages in the upper right hand corner of each grey cell represent the editing efficiency anticipated in a liver cell in each metabolic state. The ACF pre-mRNA is alternatively spliced giving rise to ACF65 and ACF 64 (shown as ACF64) and ACF45 and ACF43. ACF45/43 are nuclear and do not appear to traffick in liver cells. ACF64 trafficks between the nucleus and cytoplasm. During metabolic regulation the relative abundance of APOBEC-1, ACF64, ACF45 and ACf43 in the nucleus change (lower left hand corner). Our hypothesis is that the relative abundance of these proteins may shift the equilibrium between editosome assembly and editosome disassembly. The data leading to this hypothesis are that ACF variants have differing affinities for APOBEC-1 and apoB mRNA leading to differing ability to complement APOBEC-1 in RNA editing (upper right hand corner). For example in the ethanol treated state, net ACF nuclear import occurs, leading to APOBEC-1 nuclear accumulation. ACF64 nuclear abundance increases relative to ACF45/43 with the facilitation of ACF43, ACF64 becomes the predominant protein bound to the editing site; resulting in assembly of highly active editosomes. In contrast, during fasting, net nuclear export of ACF64 occurs and APOBEC-1 expression is reduced. ACF45 and ACF43 abundance relative to ACF64 increases promoting a competition leading to dissociation of ACF64 from apoB mRNA and from APOBEC-1. See supporting data in Sowden, M.P., Ballatori, N., De Mesy Jensen, K., Hamilton Reed, L. & Smith, H.C. Dynamic distribution of the mooring sequence RNA binding protein p66/ACF from inactive cytoplasmic complexes to active nuclear editosomes. J. Cell Biol. 115:1027-1039 (2002).
|
|